Professor Tom Owen-Hughes FRSE
Chair of Chromatin Structure and Dynamics
Molecular Cell and Developmental Biology, School of Life Sciences

Contact
Research
Chromatin remodelling and cancer epigenetics
Chromatin remodelling enzymes related to the yeast SWI/SNF complex have been found to be among the most frequently mutated tumour suppressors. Interestingly, mutations to different subunits drives cancers of different tissues. Prior, to the emergence of their importance in cancer these chromatin remodelling complexes were known to act to configure nucleosomes at key regulatory elements to enable appropriate expression of many genes. This raises the question why do ubiquitously acting transcriptional regulators act as tissue specific tumour suppressors?
Answering this question is now assisted by new tools to study acute loss of function. These include inhibitors (Papillion et al., 2018), PROTACS (Farnaby et al., 2019) and targeted protein degradation (Blümli et al., 2021). In addition to being powerful research tools, these models are of therapeutic interest as some tumours are vulnerable to loss of paralogs (Hoffman et al., 2014).
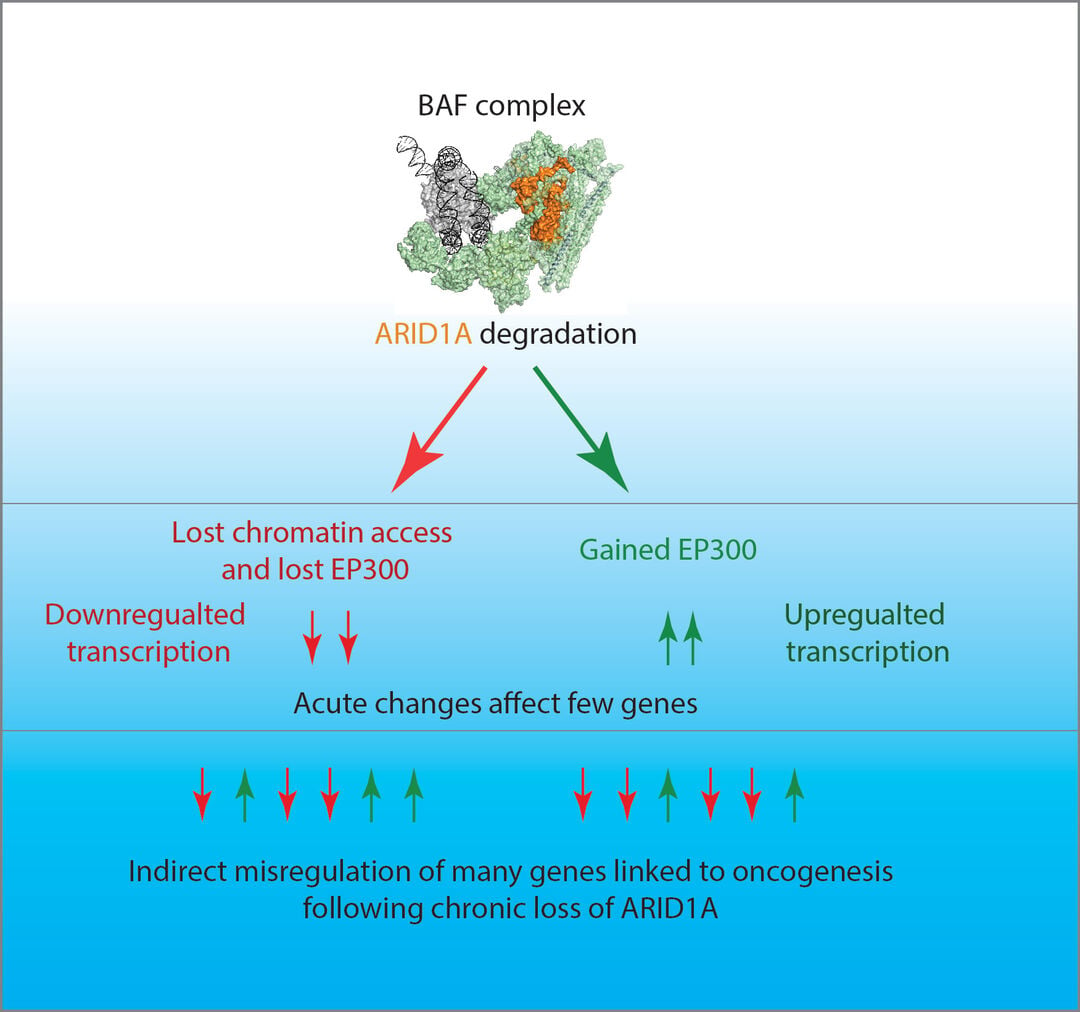
Figure 1 Acute and chronic changes following loss of ARID1A. ARID1A is a subunit of a multi-subunit complex that acts to generate accessible chromatin principally at enhancer elements (shown in orange top panel). When this subunit is degraded changes to chromatin structure, association of the coactivator EP300 and transcription are observed after just 2 hours (middle panel). Over subsequent days indirect transcriptional changes are observed at many more genes. These mimic a knock-out of the ARID1A gene. From Blümli et al., 2021.
We have used the auxin degron system to degrade subunits of SWI/SNF complexes within minutes. This has revealed that the chromatin changes occurring following loss of function affect expression of relatively few genes (Figure 1). Directly affected genes are differentially expressed after 2 hours, but in the days that follow widespread indirect changes are detected. We now reason that if we understand more about how this pathway operates, we will identify new targets that when inhibited will prevent the development of cancer.
Our ongoing work is interdisciplinary and involves engineering human stem cells for acute loss of function. We then work with our excellent stem cell facility to generate organoid models relevant to tissue specific cancers. We use high throughput DNA sequencing to characterise chromatin and transcriptional changes. We then work with computational biologists to perform integrative analysis of these time course measurements. The aim is to reveal the pathway from loss of function to development of a cancer phenotype. Longer term we aim to work with chemists to inhibit this pathway and hopefully provide new entry points for these tissue specific cancers.
Selected Publications
- Blümli S, Wiechens N, Wu MY, Singh V, Gierlinski M, Schweikert G, Gilbert N, Naughton C, Sundaramoorthy R, Varghese J, Gourlay R, Soares R, Clark D, Owen-Hughes T. Acute depletion of the ARID1A subunit of SWI/SNF complexes reveals distinct pathways for activation and repression of transcription. Cell Rep. 2021 Nov 2;37(5):109943. doi: 10.1016/j.celrep.2021.109943. PMID: 34731603; PMCID: PMC8578704.
- Sundaramoorthy R, Owen-Hughes T. Chromatin remodelling comes into focus. F1000Res. 2020 Aug 20;9:F1000 Faculty Rev-1011. doi:10.12688/f1000research.21933.1. PMID: 32864100; PMCID: PMC7445562.
- Flaus A, Downs JA, Owen-Hughes T. Histone isoforms and the oncohistone code. Curr Opin Genet Dev. 2021 Apr;67:61-66. doi: 10.1016/j.gde.2020.11.003. Epub 2020 Dec 4. PMID: 33285512.
- Farnaby W, et al., BAF complex vulnerabilities in cancer demonstrated via structure-based PROTAC design. Nat Chem Biol. 2019 Jul;15(7):672-680. doi: 10.1038/s41589-019-0294-6. Epub 2019 Jun 10. Erratum in:Nat Chem Biol. 2019 Jul 2;: PMID: 31178587; PMCID: PMC6600871.
- Sundaramoorthy R, Hughes AL, El-Mkami H, Norman DG, Ferreira H, Owen-Hughes T. Structure of the chromatin remodelling enzyme Chd1 bound to a ubiquitinylated nucleosome. Elife. 2018 Aug 6;7:e35720. doi: 10.7554/eLife.35720. PMID: 30079888; PMCID: PMC6118821.
Media availability
I am available for media commentary on my research.
Chromatin structure and gene regulation
Contact Corporate Communications for media enquiries.
Areas of expertise
- Cancer
PhD Projects
Principal supervisor
- Type
- PhD opportunity
Mechanism of action of the tumour suppressor ARID1A
Second supervisor
- Type
- PhD opportunity
Molecular basis of pioneer transcription factor function revealed by Cryo-EM
Awards
| Award | Year |
|---|---|
| Personal Fellowships / Wellcome Trust Senior Research Fellowship | 2012 |
| Fellow of the Royal Society of Edinburgh | 2009 |
| Member of the European Molecular Biology Organisation | 2007 |
| Personal Fellowships / Wellcome Trust Senior Research Fellowship | 2007 |
| National Sciences Prizes awarded since 1990 / The Colworth Medal of the British Biochemical Society | 2002 |
| International Science Prizes awarded since 1990 / EMBO Young Investigator | 2000 |
Stories
- Type
- News
School of Life Sciences hosts “The First Scottish cryo-EM symposium”
A one day symposium for the Scottish cryo-EM community took place on 30 November 2023
- Type
- News
Targeted Protein Degradation Reveals Secrets of Tumour Suppressor
PhD Student Seraina Blümli and colleagues from the Owen-Hughes lab show the ARID1A subunit of the BAF chromatin remodelling complex organised nucleosomes flanking pluripotency transcription factors and association of the coactivator EP300.
- Type
- Press release
£1.6 million award will bring machine learning into life sciences
A University of Dundee researcher has been awarded £1.6 million in funding from UK Research and Innovation (UKRI) to develop machine learning tools for the life sciences.

