Professor David Lilley FRS FRSE FRSC
Chair of Molecular Biology
Molecular Cell and Developmental Biology, School of Life Sciences

Contact
Research
Structure and function of nucleic acids
This laboratory has a general interest in the structure and dynamics of nucleic acids, protein-nucleic acid interactions and catalytic activity in RNA.
Helical branch points in DNA and their interaction with enzymes
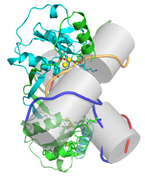
Figure 1. Resolution of a four-way DNA (Holliday) junction. Crystal structure of the DNA junction-resolving enzyme endonuclease I bound to a four-way DNA junction. J. M. Hadden et al Nature 449, 621-624 (2007).
The four-way DNA (Holliday) junction is the central intermediate of genetic recombination. The general structure of this junction was deduced over 20 years ago in Dundee. We have used single-molecule FRET methods to analyse the dynamics of the junction, demonstrating both conformer exchange and branch migration. Significant work in the group is going into the study of junction-resolving enzymes - nucleases that are selective for DNA junctions, that resolve them into their component duplex species. They exhibit exquisite structural selectivity, and a general understanding of the recognition of global DNA structure by proteins is a major goal for the laboratory.
The structure of RNA - branch points and kink turns
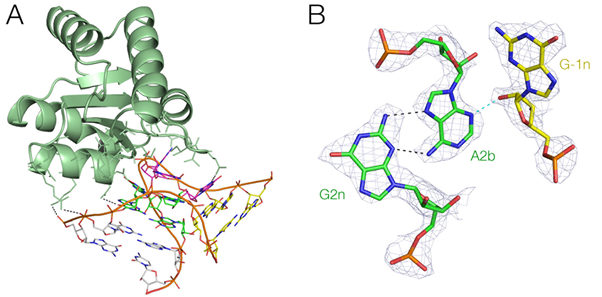
Figure 2. Protein recognition of a k-turn structure in RNA. Crystal structure of the L7Ae protein bound to a k-turn element. A. The complex. B. A key A-minor triple interaction in the k-turn, with the electron density from the composite, omit map shown contoured at 2-sigma. L. Huang and D. M. J. Lilley RNA In the press.
Helical branch points are key elements that determine the architecture of large RNA assemblies as well as smaller, autonomously-folding RNA species. These include three- and four-way helical junctions as found in various ribozymes and IRES elements. k-turns are ubiquitous elements in RNA, that generate a tight kink in the helical axis. The act to mediate tertiary interactions, and to create specific protein binding sites. They are important in translation (many examples in the ribosome), RNA modification (box C/D a H/ACA snoRNA), spliceosome assembly (U4 snRNA) and riboswitches (7 examples now known). Our goal is to understand the connection between sequence, structure and function, using a combination of X-ray crystallography and single-molecule FRET.
Catalytic activity in RNA - mechanism of ribozyme action
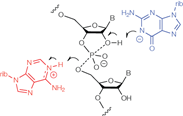
Figure 3. The catalytic mechanism of the hairpin and VS ribozymes. In the cleavage reaction, both ribozymes use a guanine nucleobase (blue) as the general base to deprotonate the 2'-hydroxyl group, and an adenine nucleobase (red) as the general acid to protonate the O2' oxyanion leaving group. T. J. Wilson et al Proc. Natl. Acad. Sci. USA 107, 11751–11756 (2010).
Ribozymes are enzymes that are made of RNA rather than protein. A major goal is to understand how RNA can accelerate chemical reactions by a million-fold or more, with a fraction of the chemical resources of a protein. We are studying the mechanism of the nucleolytic ribozymes, and have provided good evidence for nucleobase-mediated general acid-base catalysis. For example, we have shown that both the Varkud satellite and hairpin ribozymes act by using guanine and adenine nucleobases as base and acid respectively in their cleavage reactions.
Fluorescence resonance energy transfer as a tool in the structural biology of nucleic acids
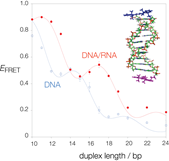
Figure 4. Orientation dependence in FRET. Plot of FRET efficiency (EFRET) as a function of duplex length for a series of DNA (blue) and DNA/RNA hybrid (red) duplexes 5'-terminally labelled with Cy3 and Cy5 fluorophores (inset). The modulation is due to the parameter kappa-squared, which measures the relative orientation between the transition dipole moments. Note the shift between the two series, arising from the different helical geometries of the B- and A-form helices of DNA and DNA/RNA respectively. A. Iqbal et al Proc. Natl. Acad. Sci. USA 105, 11176-11181 (2008).
FRET is a sensitive spectroscopic method to study the conformation of macromolecules. It arises from dipolar interaction between the transition dipole moments of two different fluorophores and depends on both their separation and relative orientation. We have shown that the orientational parameter kappa-squared is not simply a complication in the extraction of reliable distance information, but a valuable source of structural data. We have shown that we can use FRET between indocarbocyanine fluorophores terminally attached to DNA or RNA to provide orientational information.
Selected Publications
Selected from a total of 354 papers (current h index = 73)
- T. J. Wilson and D. M. J. Lilley The evolution of chemistry in the ribozymes Science 323, 1436-1438 (2009).
- T. J. Wilson, N.-S. Li, J. Lu, J. K. Frederiksen, J. A. Piccirilli and D. M. J. Lilley Nucleobase-mediated general acid-base catalysis in the Varkud satellite ribozyme Proc. Natl. Acad. Sci. USA 107, 11751–11756 (2010).
- K. T. Schroeder, P. Daldrop and D. M. J. Lilley. RNA tertiary interactions in a riboswitch stabilize the structure of a kink turn. Structure 19, 1233-1240 (2011).
- J. Ouellet, S. Schorr, A. Iqbal, T. J. Wilson, and D. M. J. Lilley Orientation of cyanine fluorophores terminally attached to DNA via long, flexible tethers. Biophys. J. 101, 1148-1154 (2011).
- S. Kath-Schorr, T. J. Wilson, N.-S. Li, J. Lu, J. A. Piccirilli and D. M. J. Lilley General acid-base catalysis mediated by nucleobases in the hairpin ribozyme. J. Amer. Chem. Soc. 134, 16717–16724 (2012).
- J. Wang, T. Fessl, K. T. Schroeder, J. Ouellet, Y. Liu, A. D. J. Freeman and D. M. J. Lilley Single-molecule observation of the induction of k-turn RNA structure on binding L7Ae protein. Biophysical. J. 103, 2541-2548 (2012).
Teaching
Courses on nucleic acid structure and function, enzyme mechanism and fluorescence
Media availability
I am available for media commentary on my research.
Nucleic acid structure
Contact Corporate Communications for media enquiries.
Areas of expertise
- Cancer
Awards
| Award | Year |
|---|---|
| National Sciences Prizes awarded since 1990 / Royal Society of Chemistry Khorana Prize | 2016 |
| National Sciences Prizes awarded since 1990 / RNA Society Lifetime Service Award | 2015 |
| National Sciences Prizes awarded since 1990 / Royal Society of Chemistry, Interdisciplinary Award | 2006 |
| Fellow of the Royal Society | 2002 |
| National Sciences Prizes awarded since 1990 / RNA and Ribozyme Chemistry Award of The Royal Society of Chemistry | 2002 |
| International Science Prizes awarded since 1990 / The Prelog Medal in Stereochemistry, Zurich, Switzerland | 1996 |
| International Science Prizes awarded since 1990 / The Gold Medal of G. Mendel of the Czech Academy of Sciences | 1994 |
| Fellow of the Royal Society of Edinburgh | 1988 |
| Member of the European Molecular Biology Organisation | 1984 |
Stories
- Type
- News
Artificial intelligence in life sciences and medicine – conference in Tianjin, China
Professor David Lilley addressed 500 people at a conference on application of supercomputing at Nankai university in Tianjin, PR China.
- Type
- News
A chemical glimpse into life’s origins
How did life start out on this planet, around 3.5 billion years ago? Since we cannot go back in time to look we can only get hints by projecting backwards from contemporary life, and doing experiments that test the limits of what might have been possible.
- Type
- News
David Lilley made visiting professor at Nankai University
Professor David Lilley has been made a visiting professor at Nankai University in Tianjin, China. Nankai is one of the top 20 universities in China and is celebrating its 100th Anniversary this year.
