Dr Wataru Kobayashi
Principal Investigator
Life Sciences Office, School of Life Sciences
Molecular Cell and Developmental Biology, School of Life Sciences

Contact
Biography
Wataru Kobayashi is a Principal Investigator within the Division of Molecular Cell & Developmental Biology. He studied biochemistry and structural biology with a focus on understanding how nucleosome structure regulates biological processes in the cell. In 2016, he obtained his PhD from Waseda University under the supervision of Prof Hitoshi Kurumizaka. He continued as a Research Associate at Waseda University (2016-2019).
From 2019 to 2025, Wataru was a post-doctoral researcher at the Institute of Molecular Biotechnology (IMBA) and the Max Planck Institute of Biochemistry in the group of Prof Kikuë Tachibana. During this time, he expanded his expertise in embryology and genomics, investigating the molecular mechanisms underlying zygotic genome activation.
Wataru joined the University of Dundee to establish his own group, which focuses on the molecular mechanisms by which transcription factors regulate epigenetic reprogramming in early mammalian development. In 2026, he obtained a Wellcome Career Development Award.
Research
The life of a mammal begins with a single cell, the fertilized egg. During mammalian pre-implantation development, embryos are initially awakened during zygotic genome activation and further develop into three distinct cell lineages at the blastocyst stage. The outer layer of the blastocyst consists of trophectoderm cells, which form extraembryonic tissues such as the placenta. The inner cell mass differentiates into epiblast cells and primitive endoderm cells, which give rise to the embryo proper and yolk sac, respectively. Although most cells contain the same genomic information, they differentiate into a wide variety of distinct cell types and develop into a complex multicellular organism.
How do cells achieve such drastic changes in cell fate during early development? Our laboratory is interested in how embryos control transcriptional and epigenetic regulation at the beginning of life.
Transcription factors play a key role in establishing gene regulatory networks that determine cell fate. Notably, specialized transcription factors, called pioneer transcription factors, are uniquely capable of engaging closed chromatin and modulating epigenetic and chromatin states by recruiting chromatin remodelers and histone modifiers. These factors are key drivers in development and cell differentiation. However, the molecular mechanisms by which transcription factors regulate gene regulatory networks and epigenome during early development are largely unknown, in large part due to the scarcity of mammalian embryo samples.
To address this fundamental question, we use genomics methods that generate robust data from low-input materials. By combining this with in vitro reconstitution and structural analysis using cryo-electron microscopy, we aim to understand the molecular mechanisms of mammalian pre-implantation development across scales from atoms to embryo.
With 1 in 6 people worldwide affected by infertility, as reported by the WHO, understanding the molecular underpinnings of early embryonic failure and pregnancy loss is one of the most urgent challenges in reproductive biology. Our long-term goal is to translate principles from the mouse model to human development, with broad clinical implications for understanding infertility and miscarriage.
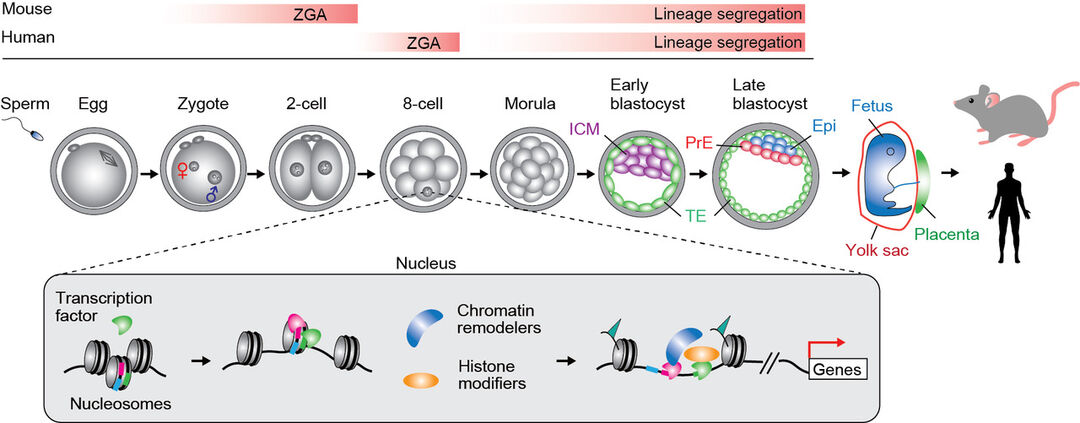
Research overview: A schematic illustration showing mammalian early development. Abbreviations: EPI, epiblast; ICM, inner cell mass; PrE, primitive endoderm; TE, trophectoderm; ZGA, zygotic genome activation.
Key publications
For a full list of publications, see Google Scholar.
* = co-first author
- Kobayashi W*, Ruangroengkulrith S*, Arslantas E, Mohanan A, Tachibana K. Feed-forward loops by NR5A2 ensure robust gene activation during pre-implantation development, Development. 153:dev205059 (2026).
- Kobayashi W †, Michael AK †, Ruangroengkulrith S, Kümmecke M, Tachibana K †. Protocol for integrative analysis of transcription factor-nucleosome interactions using SeEN-seq and cryo-EM structure determination. STAR Protoc. 7:104295 (2025).
- Kobayashi W, Tachibana K. CUT&Tag for transcription factor binding profiles during mammalian zygotic genome activation. Methods in Molecular Biology. 2923, 77-87 (2025).
- Kobayashi W, Sappler H. A, Bollschweiler D, Kümmecke M, Basquin J, Arslantas E, Ruangroengkulrith S, Hornberger R, Duderstadt K, Tachibana K. Nucleosome-bound NR5A2 structure reveals pioneer factor mechanism by minor groove anchor competition. Nat. Struct Mol. Biol., 31, 757-766 (2024).
- Gassler J*, Kobayashi W*, Gáspár I*, Ruangroengkulrith S*, Mohanan A, Gómez Hernández L, Kravchenko P, Kümmecke M, Lalic A, Rifel N, Ashburn RJ, Zaczek M, Vallot A, Cuenca Rico L, Ladstätter S, Tachibana K. Zygotic genome activation by the totipotency pioneer factor Nr5a2. Science 378, 1305-1315 (2022).
- Kobayashi W, Takizawa Y, Aihara M, Negishi L, Ishii H, Kurumizaka H. Structural and biochemical analyses of the nuclear pore complex component ELYS identify residues responsible for nucleosome binding. Commun Biol. 163 (2019)
PhD Projects
Principal supervisor
- Type
- PhD opportunity
Molecular basis of pioneer transcription factor function revealed by Cryo-EM
Awards
| Award | Year |
|---|---|
| Major Personal Funding Awards / Wellcome Career Development Award | 2026 |
Stories
- Type
- News
Wellcome Career Development Awards awarded to Dr Wataru Kobayashi
Dr Wataru Kobayashi has been awarded £ 2.16M from the Wellcome Trust to study the earliest step of mammalian development
