
Contact
Biography
I graduated from the University of Liverpool with a first degree in Marine Biology. I studied cyanobacteria for my PhD at the University of Dundee with Pete Rowell and Sir William Stewart. I first worked on RNA with John Brown in collaboration with Jean Beggs from the University of Edinburgh. I won a Royal Society fellowship to join Witold Filipowicz’s lab at the Friedrich Miescher Institute, Basel, Switzerland and later joined Dame Caroline Dean’s lab at the John Innes Centre. I returned to the University of Dundee as a Lecturer, Reader, and then Professor of Molecular Genetics.
Research
RNA processing and modification
We study RNA. Although the genes in our chromosomes are made of DNA, they are transcribed into copies of RNA when switched on. RNA is processed and modified in ways that ultimately determine what a gene codes for. We study processing events called splicing, polyadenylation, and RNA modification. In this way, we learn what genomes encode and how the regulation of genes leads to changes in biology.
Our science is curiosity-led and attempts to answer basic questions about RNA and biology. We try to use unbiased approaches that include forward genetic screens, global sequencing of messenger RNAs and inter-species association mapping. We combine expertise in RNA molecular biology with genetics and computational biology.
Our most important recent discovery is uncovering two classes of 5’ splice sites in eukaryotic introns. We can separate these classes in the genomes of diverse species and reveal genetic backgrounds that sensitise them. We have demonstrated that the two splice site classes are associated with splicing complexity. Recently identified genetic variants that cause the neurodevelopmental disorder ReNu syndrome sensitise the same two classes we first found depended on U6 snRNA m6A modification. The two classes likely reflect the importance of driving the transfer of 5’ splice sites to U5 and U6 snRNAs for selection. In summary, two classes of 5’ splice sites are a fundamental but overlooked feature of eukaryotic genomes.
We try to be as transparent as possible in our scientific process. We freely share all our source data and the code we use to analyse our data sets. We pre-print all our work and publish it in open-access scientific journals with transparent peer review so anyone can read it for free. Our public engagement activities include a genetics garden we established at the University of Dundee Botanic Garden more than 10 years ago.
Our work is publicly funded by the UK government through BBSRC and GCRF, and by the EU through Horizon 2020.
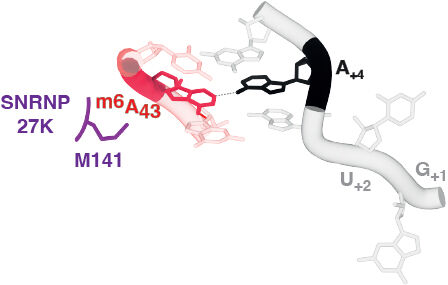
Figure 1. Disrupting m6A modification of the U6 snRNA ACAGA box or Methionine 141 of SNRNP27K sensitises two splice site classes. Recent cryo-EM analysis of spliceosomes from Reinhard Lührmann’s lab reveals how closely connected these interactions are – in time and space - during 5’ splice site selection (figure prepared by our collaborator Dr. Sebastian Fica, Department of Biochemistry, University of Oxford).
Selected Publications
Metheringham, C.L., Srivastava, A.K., Thorpe, P., Maji, A., Parker, M.T., Barton, G.J., Simpson, G.G. (2025). Disruption of the mRNA m6A writer complex triggers autoimmunity in Arabidopsis. In bioRxiv 2025.02.18.638636. https://doi.org/10.1101/2025.02.18.638636
Parker, M.T., Fica, S.M. & Simpson, G.G. (2025) RNA Splicing: A split consensus reveals two major 5’ splice site classes. Open Biol 15: 240293 https://royalsocietypublishing.org/rsob/article/15/1/240293/91501/RNA-splicing-a-split-consensus-reveals-two-major-5
Shen, A., Hencel, K., Parker, M.T., Scott, R.,Skukan, R., Adesina, A.S., Metheringham, C.L., Miska, E.A., Nam, Y., Haerty, W., Simpson, G.G., Akay, A. (2024) U6 snRNA m6A modification is required for accurate and efficient splicing of C. elegans and human pre-mRNAs. Nucleic Acids Research 2024 gkae 447 https://doi.org/10.1093/nar/gkae447
Parker, M. T., Fica, S. M., Barton, G. J. & Simpson, G. G. (2023) Inter-species association mapping links splice site evolution to METTL16 and SNRNP27K. eLife, 12, e91997. https://elifesciences.org/articles/91997
Parker, M. T., Soanes, B. K., Kusakina, J., Larrieu, A., Knop, K., Joy, N., Breidenbach, F., Sherwood, A. V., Barton, G. J., Fica, S. M., Davies, B. H., & Simpson, G. G. (2022). m6A modification of U6 snRNA modulates usage of two major classes of pre-mRNA 5’ splice site. eLife, 11, e78808. https://elifesciences.org/articles/78808
Zhang, M., Bodi, Z., Mackinnon, K., Zhong, S., Archer, N., Mongan, N. P., Simpson, G. G., & Fray, R. G. (2022). Two zinc finger proteins with functions in m6A writing interact with HAKAI. Nature Communications, 13(1), 1127. https://doi.org/10.1038/s41467-022-28753-3
Bredeson, J.V., Lyons, J.B., Oniyinde, I.O., Okereke, N.R., Kolade, 0., Nnabue, I., Nwadili, C.O., Hřibová,E., parker, M.T.,Nwogha,J., Shu, S., Carlson, J., Kariba, R., Muthemba, S., knop, K., Barton, G.J., Sherwood, A.V., Lopez-Montes, A., Asiedu, R., Jamnadass, R., Muchugi, A., Goodstein, D., Egesi, C.N., Featherston, J., Asfaw, A., Simpson, G.G., Doležel, J., Hendre, P.S., Van Deynze, A., Kumar, P.L., Obidiegwu, J.E., Bhattacharjee, R., Rokhsar, D.S. (2022). Chromosome evolution and the genetic basis of agronomically important traits in greater yam. Nature Communications, 13(1), 2001. https://doi.org/10.1038/s41467-022-29114-w
Parker, M. T., Barton, G. J., & Simpson, G. G. (2021). Yanocomp: robust prediction of m6A modifications in individual nanopore direct RNA reads. In bioRxiv (p. 2021.06.15.448494). https://doi.org/10.1101/2021.06.15.448494
Parker, M. T., Knop, K., Zacharaki, V., Sherwood, A. V., Tomé, D., Yu, X., Martin, P. G., Beynon, J., Michaels, S. D., Barton, G. J., & Simpson, G. G. (2021). Widespread premature transcription termination of Arabidopsis thaliana NLR genes by the spen protein FPA. eLife, 10, e65537. https://doi.org/10.7554/eLife.65537
Parker, M. T., Knop, K., Barton, G. J., & Simpson, G. G. (2021). 2passtools: two-pass alignment using machine-learning-filtered splice junctions increases the accuracy of intron detection in long-read RNA sequencing. In Genome Biology 22, 72. https://doi.org/10.1186/s13059-021-02296-0
Parker, M. T., Knop, K., Sherwood, A. V., Schurch, N. J., Mackinnon, K., Gould, P. D., Hall, A. J., Barton, G. J., & Simpson, G. G. (2020). Nanopore direct RNA sequencing maps the complexity of Arabidopsis mRNA processing and m6A modification. eLife, 9, e49658. https://doi.org/10.7554/eLife.49658