Press release
Covid virus images shared for global science community
Scientists at the University of Dundee and the EMBL’s European Bioinformatics Institute (EMBL-EBI) have published online some of the largest and highest resolution images yet recorded of the SARS-CoV-2 virus, the cause of the Covid-19 pandemic.
Published on 1 May 2020
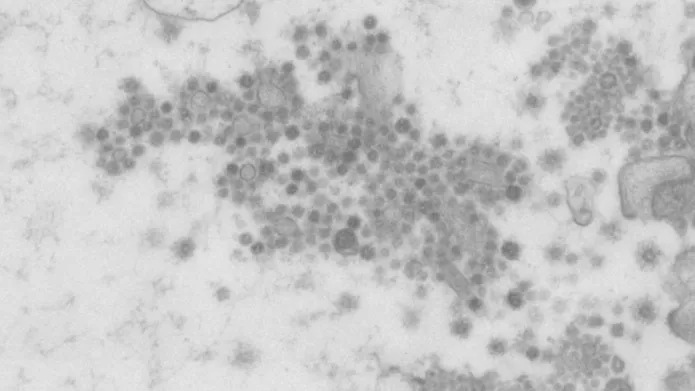
Image shows part of the wider view of SARS-CoV-2 in human intestinal cells
Scientists at the University of Dundee and the EMBL’s European Bioinformatics Institute (EMBL-EBI) have published online some of the largest and highest resolution images yet recorded of the SARS-CoV-2 virus, the cause of the Covid-19 pandemic.
The images were collected by a consortium of researchers from the Hubrecht Institute in Utrecht, Erasmus MC University Medical Center Rotterdam, and Maastricht University in the Netherlands.
They show the formation of SARS-CoV-2 particles, the virus at the heart of the current COVID-19 pandemic, in a tissue model of the human gut as viewed in an ultra-powerful electron microscope, which may be a significant breakthrough in understanding how the disease can infect the intestine.
Each of the images is greater than 30-50 GBytes – which is 500 to 1000 times larger than an image recorded on an iPhone-- and the scientists who collected them were only able to include very small parts of the images in their paper, published today (May 1, 2020) in the journal Science.
Since the images and data are the first of their kind ever recorded of SARS-CoV-2, it is important they can be widely shared with the global scientific community. To share these massive chunks of unique data, the Dutch team turned to a group of computer scientists at the University of Dundee's School of Life Sciences, led by Professor Jason Swedlow, who have built and run the Image Data Resource (IDR), a publication system for very large, complex images recorded using the world's most powerful microscopes and imaging systems.
IDR uses the latest data handling and cloud computing technology to make these very large, very valuable datasets publicly viewable and available for analysis and download.
There are two datasets available in the IDR – ( Image 1 and Image 2) – which show the virus assembling and leaving the human intestinal cells.
The data files generated by Maastricht University scientists showing SARS-CoV-2 locations in the model of the human gut are also being deposited in the EMBL-EBI’s EMPIAR archive for wider distribution to the biomedical community (Datasets). Scientists can download the original data and re-analyse them using their own tools and methods.
Professor Swedlow said, “We're excited to publish these important new datasets in IDR, where they can be seen by researchers around the world, who can also scan the images and view the SARS-CoV-2 virus up close on their computer.
“We have included annotations from the authors so anyone who reads the paper from the research teams in the Netherlands can easily see what the authors published, but also can examine other parts of the data and maybe make their own discoveries.
“This kind of sharing of data has never been more important than in our current situation where we urgently need to work together around the world to find out more about this disease and ultimately be able to treat or control it.”
The IDR is a cloud-based image data publication system built and run by Professor Swedlow's Open Microscopy Environment group at the University of Dundee, in collaboration with Dr Alvis Brazma and Dr Ugis Sarkans at EMBL's European Bioinformatics Institute. It has published over 170 TBytes of highly valuable imaging data from over 70 studies.
Dr Ardan Patwardhan, who heads up the EMPIAR archive, said, “These images of SARS-CoV-2 virus particles in the context of COVID-19 patient tissue are of enormous value for the scientific community. The authors are to be commended for taking the initiative to make their data publicly available and we hope this will inspire others to follow suit. We are working with the authors and Professor Swedlow’s team to make the electron microscopy data available via EMPIAR and linked to IDR.”
Dr Frances Wong, the Curator on Professor Swedlow's IDR team said, “It has been an honour to work with the data from Dr Clevers' and Dr Ravelli's groups in the Netherlands. The images are amazing, and I hope the additional annotations our team has added increases the utility of the data for the global scientific community."
IDR was built and is run through funding from the UKRI's Biotechnology and Biological Sciences Research Council and the Wellcome Trust. EMPIAR was built and is run through funding from UKRI’s Medical Research Council and the Wellcome Trust. IDR and EMPIAR are part of an ecosystem of imaging resources being constructed by EMBL-EBI that includes its recently announced BioImage Archive, funded through UKRI’s Strategic Priority Fund.
The research from the Hubrecht Institute in Utrecht, Erasmus MC University Medical Center Rotterdam, and Maastricht University, has found that the coronavirus SARS-CoV-2, which causes COVID-19, can infect the cells of the intestine and multiply there.
Using state-of-the-art cell culture models of the human intestine, the researchers have successfully propagated the virus in vitro, and monitored the response of the cells to the virus, providing a new cell culture model for the study of COVID-19. These findings could explain the observation that approximately one third of COVID-19 patients experience gastrointestinal symptoms such as diarrhoea, and the fact that the virus often can be detected in stool samples.
Patients with COVID-19 show a variety of symptoms associated with respiratory organs - such as coughing, sneezing, shortness of breath, and fever - and the disease is transmitted via tiny droplets that are spread mainly through coughing and sneezing. One third of the patients however also have gastrointestinal symptoms, such as nausea and diarrhoea. In addition, the virus can be detected in human stool long after the respiratory symptoms have been resolved. This suggests that the virus can also spread via so-called “fecal-oral transmission”.
Publication SARS-CoV-2 productively Infects Human Gut Enterocytes Mart M. Lamers*, Joep Beumer*, Jelte van der Vaart*, Kèvin Knoops, Jens Puschhof, Tim I. Breugem, Raimond B.G. Ravelli, J. Paul van Schayck, Anna Z. Mykytyn, Hans Q. Duimel, Elly van Donselaar, Samra Riesebosch, Helma J.H. Kuijpers, Debby Schipper, Willine J. van de Wetering, Miranda de Graaf, Marion Koopmans, Edwin Cuppen, Peter J. Peters, Bart L. Haagmans† and Hans Clevers†. Science 2020.
* Equal contribution, † equal contribution.
This study was a collaboration between the Hubrecht Institute in Utrecht, the Erasmus MC University Medical Center Rotterdam, Maastricht University, the UMC Utrecht and Single Cell Discoveries in the Netherlands.
The microscopy data are publicly available via the Image Data Resource (idr0083, https://idr.openmicroscopy.org ) and the EMPIAR Archive (10404, https://www.ebi.ac.uk/pdbe/emdb/empiar/ ) - with help from the University of Dundee and the European Bioinformatics Institute) and the genomic data are publicly available via the Gene Expression Omnibus (GSE149312, https://www.ncbi.nlm.nih.gov/geo ), to ensure efficient sharing of data related to COVID-19 between researchers all across the world.
European Bioinformatics Institute (EMBL-EBI)
The European Bioinformatics Institute (EMBL-EBI) is a global leader in the storage, analysis and dissemination of large biological datasets. We help scientists realise the potential of 'big data' by enhancing their ability to exploit complex information to make discoveries that benefit humankind.
We are at the forefront of computational biology research, with work spanning sequence analysis methods, multi-dimensional statistical analysis and data-driven biological discovery, from plant biology to mammalian development and disease.
We are part of EMBL and are located on the Wellcome Genome Campus, one of the world's largest concentrations of scientific and technical expertise in genomics.
Website: www.ebi.ac.uk
Open Microscopy Environment (OME) at the University of Dundee
OME produces open tools to support data management for microscopy. Designed to interact with open and commercial software, all OME formats and software are free, and all OME source code is available under open source licenses. OME’s Bio-Formats library is a global standard for reading and accessing bioimaging data. OME’s OMERO is used in 1000s of labs and institutions for managing, sharing and publishing these data. OME is based at the School of Life Sciences, University of Dundee and run by Prof Jason Swedlow. Since 2015, OME has been collaborating with EMBL-EBI to build and run the Image Data Resource (IDR), one of world’s largest collections of public bioimage data and metadata.
Website: www.openmicroscopy.org